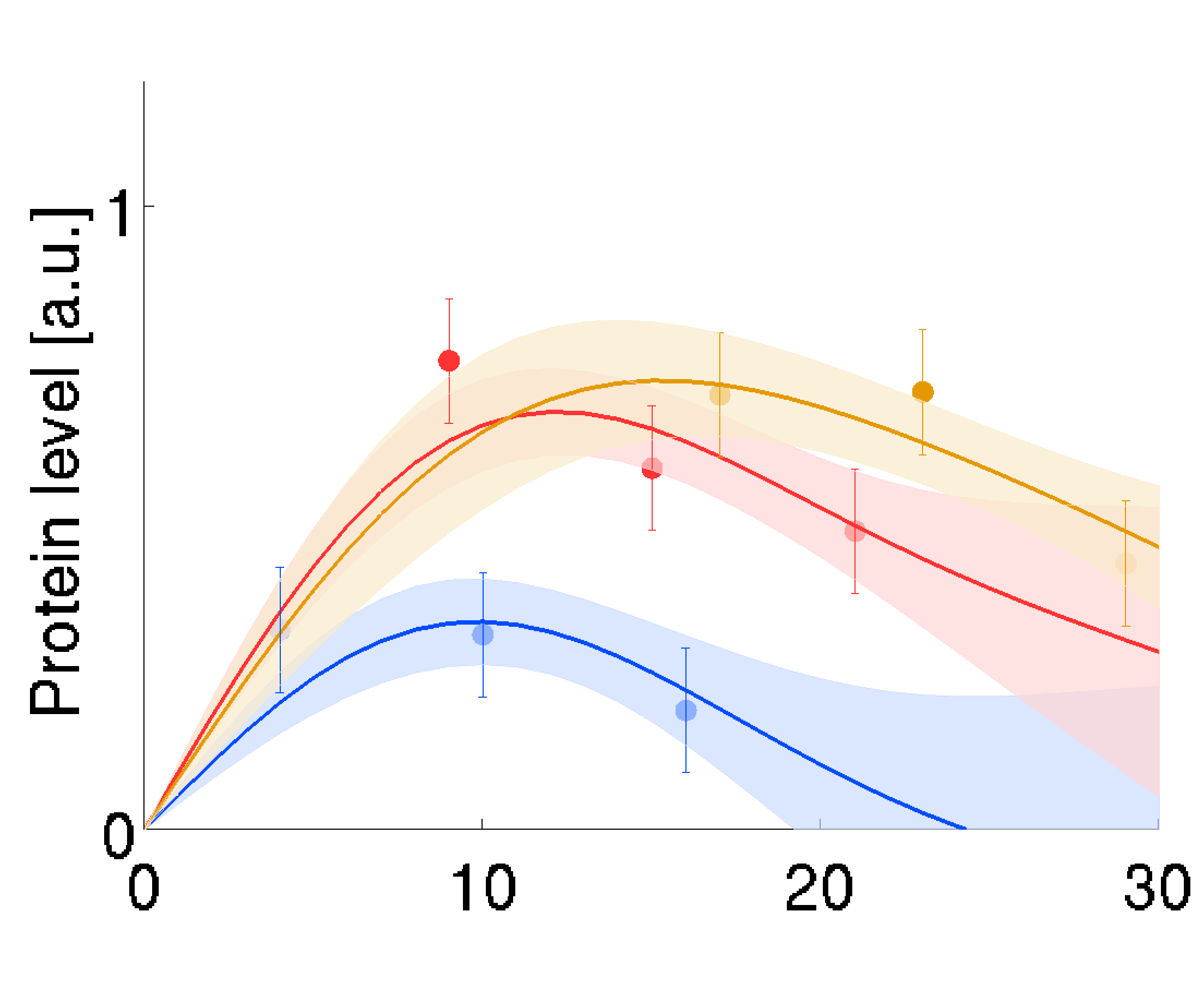
Proteomics time series measured in vivo show quantification errors and biological variability. Disentangling these effects from each other can be a challenge. We developed replicate regression specifically for proteomics and other omics time series measured in replicate. It dissects quantification errors from biological variability and fits the replicate time series by similar, individual regression curves. Assuming a linear degradation or dilution of proteins, we can directly estimate time-dependent rates of protein production. The regression is based on Bayesian estimation and yields an ensemble of most plausible curves, which are summarised by mean curves and uncertainty ranges. Prior distributions determine how smooth and similar the regression curves will be. These priors can be manually chosen or be learnt from large omics data sets.